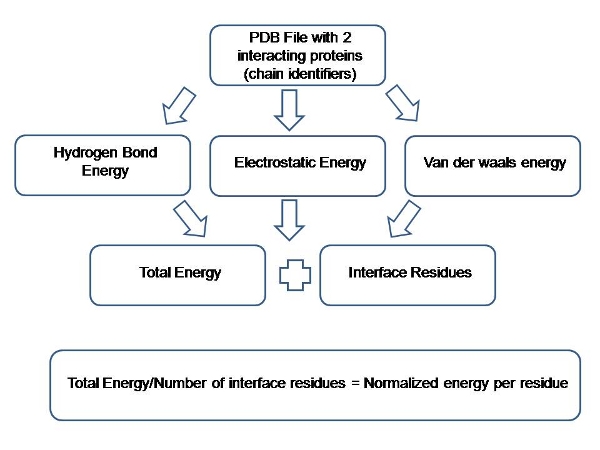
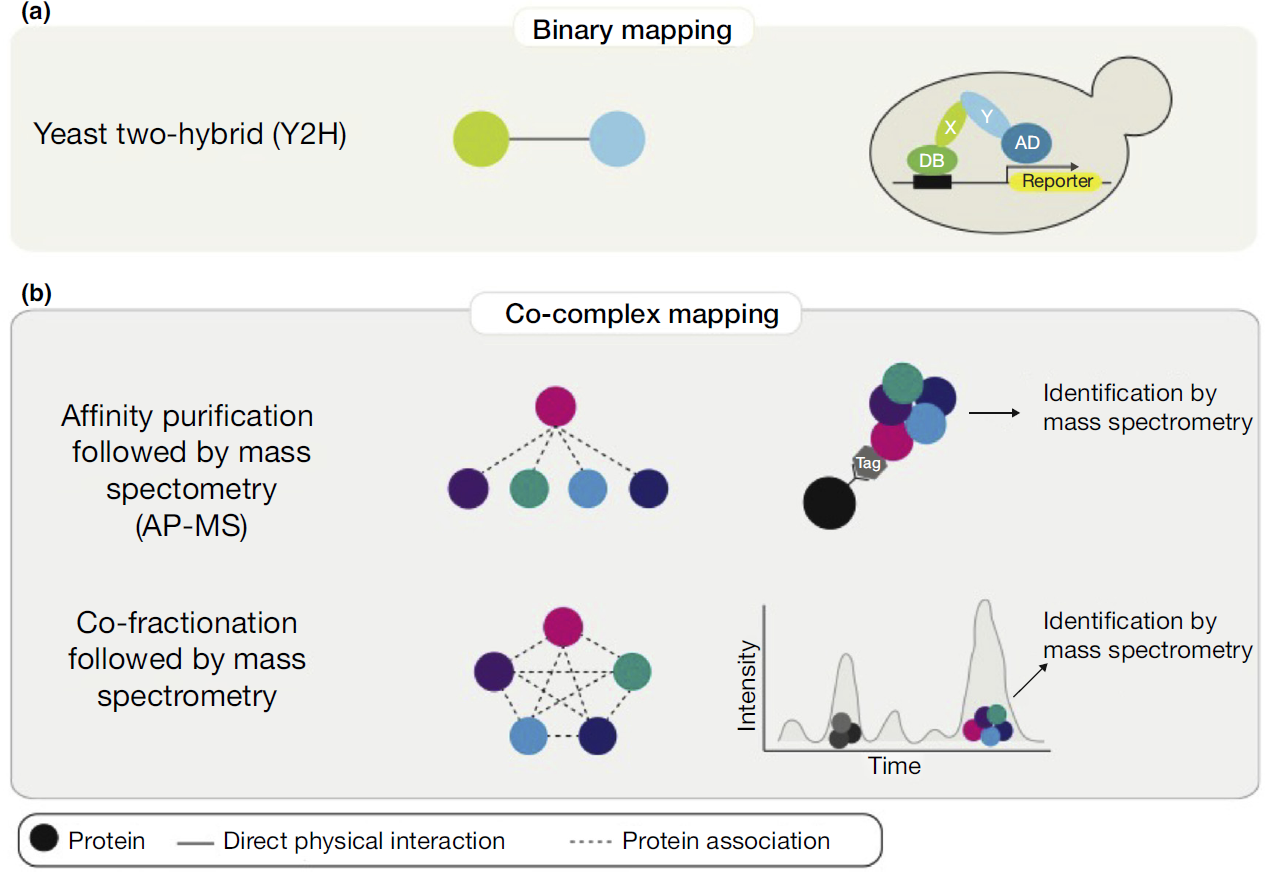
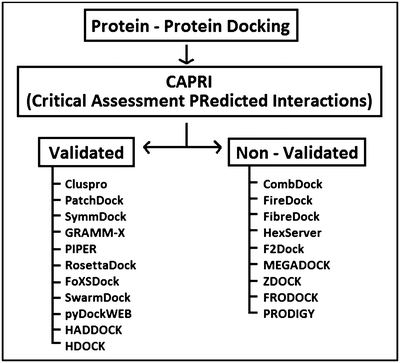
New SDC function prediction based on protein-protein interaction using bioinformatics tools - ScienceDirect

Tools for protein interaction analyses that can also be useful in PTM... | Download Scientific Diagram

BindML: Software for Predicting Protein-Protein Interaction Sites using Phylogenetic Substitution Models

Protein−protein interaction network analysis. Networks, constructed by... | Download Scientific Diagram

New methods to modulate protein-protein interactions for drug discovery : Andrew Wilson Research Group
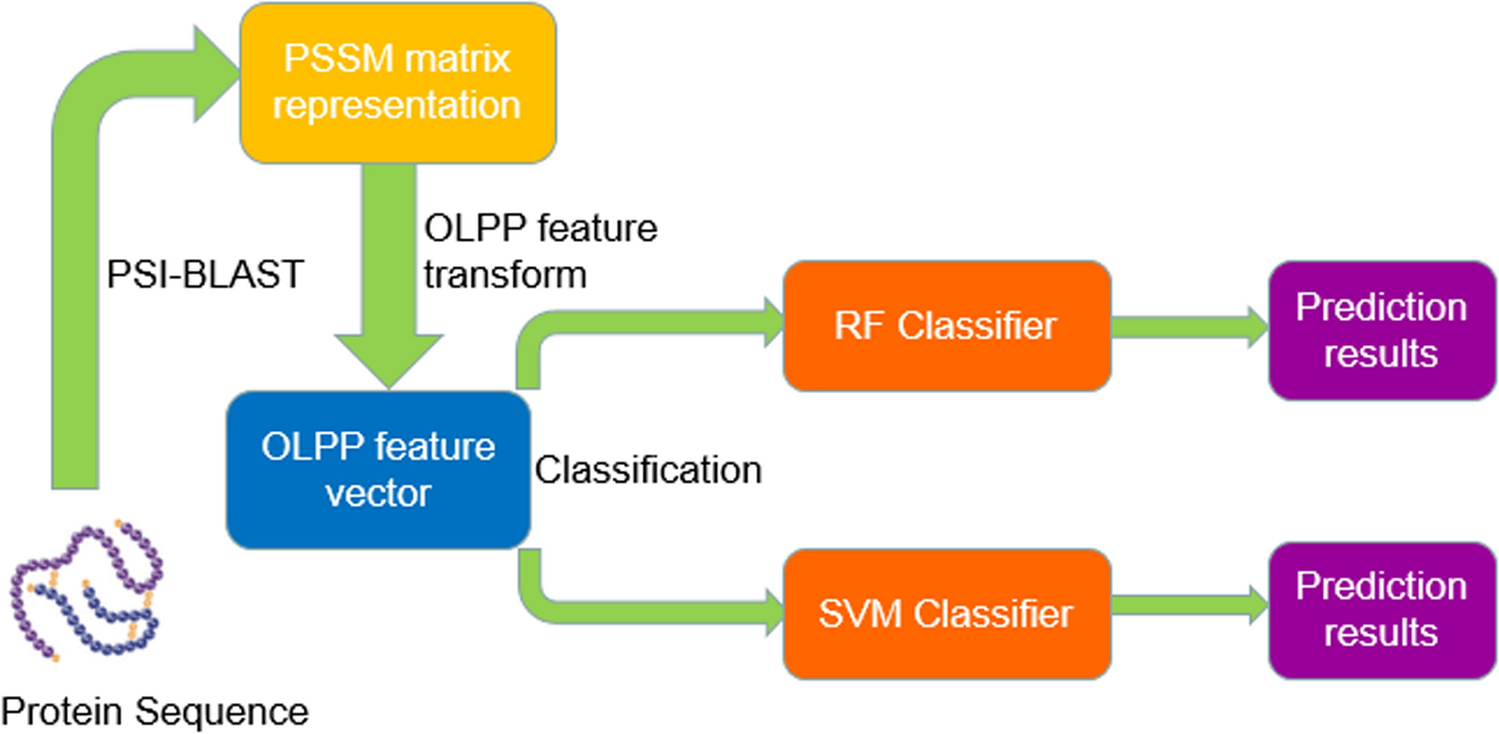
Robust and accurate prediction of protein–protein interactions by exploiting evolutionary information | Scientific Reports
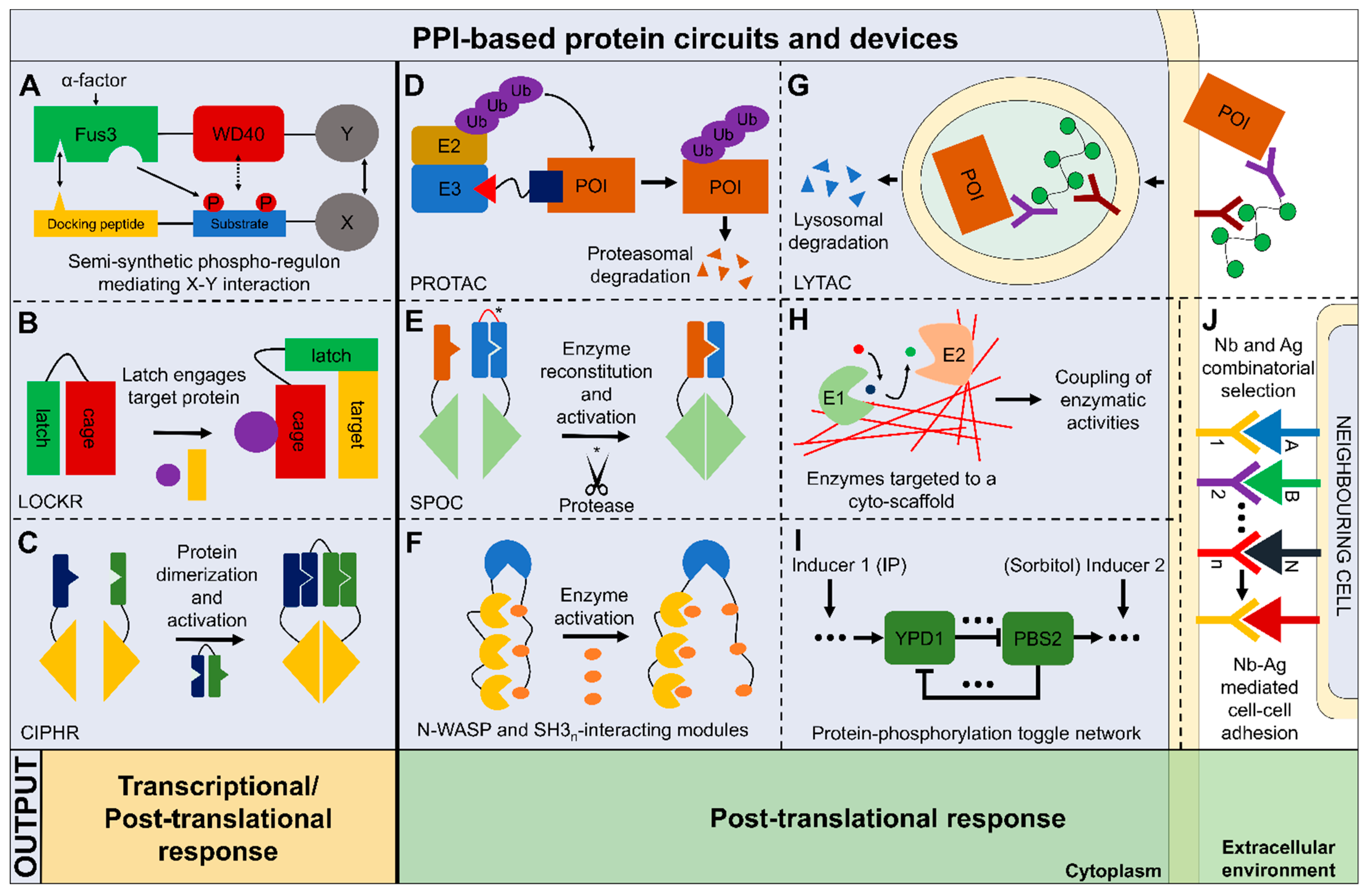
Life | Free Full-Text | Synthetic Protein Circuits and Devices Based on Reversible Protein-Protein Interactions: An Overview

New Tools Illuminate Protein-Protein Interactions: Innovative methods and products allow more detailed studies of cellular dynamics: Genetic Engineering & Biotechnology News: Vol 41, No 8
![PDF] TRI_tool: a web-tool for prediction of protein–protein interactions in human transcriptional regulation | Semantic Scholar PDF] TRI_tool: a web-tool for prediction of protein–protein interactions in human transcriptional regulation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e6a5a4d82b658dabb2a2cc770b98f8600d763eb6/2-Figure1-1.png)
PDF] TRI_tool: a web-tool for prediction of protein–protein interactions in human transcriptional regulation | Semantic Scholar
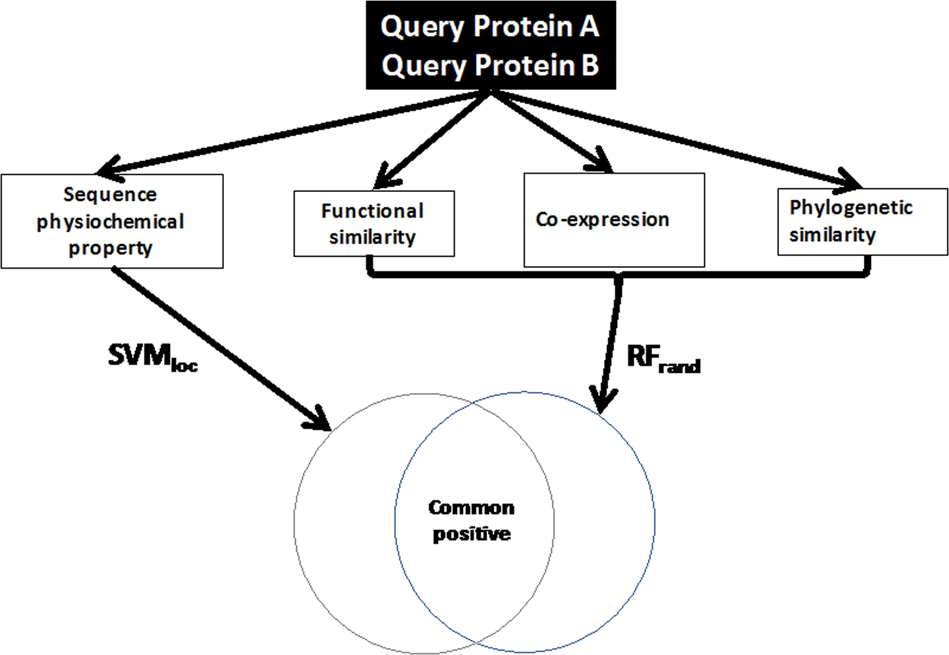
Computational identification of protein-protein interactions in model plant proteomes | Scientific Reports

Amazon.com: Protein-Protein Interactions - Computational and Experimental Tools: 9789535103974: Cai, Weibo, Hong, Hao: Books

Building Protein-Protein Interaction Networks with Proteomics and Informatics Tools* | Semantic Scholar
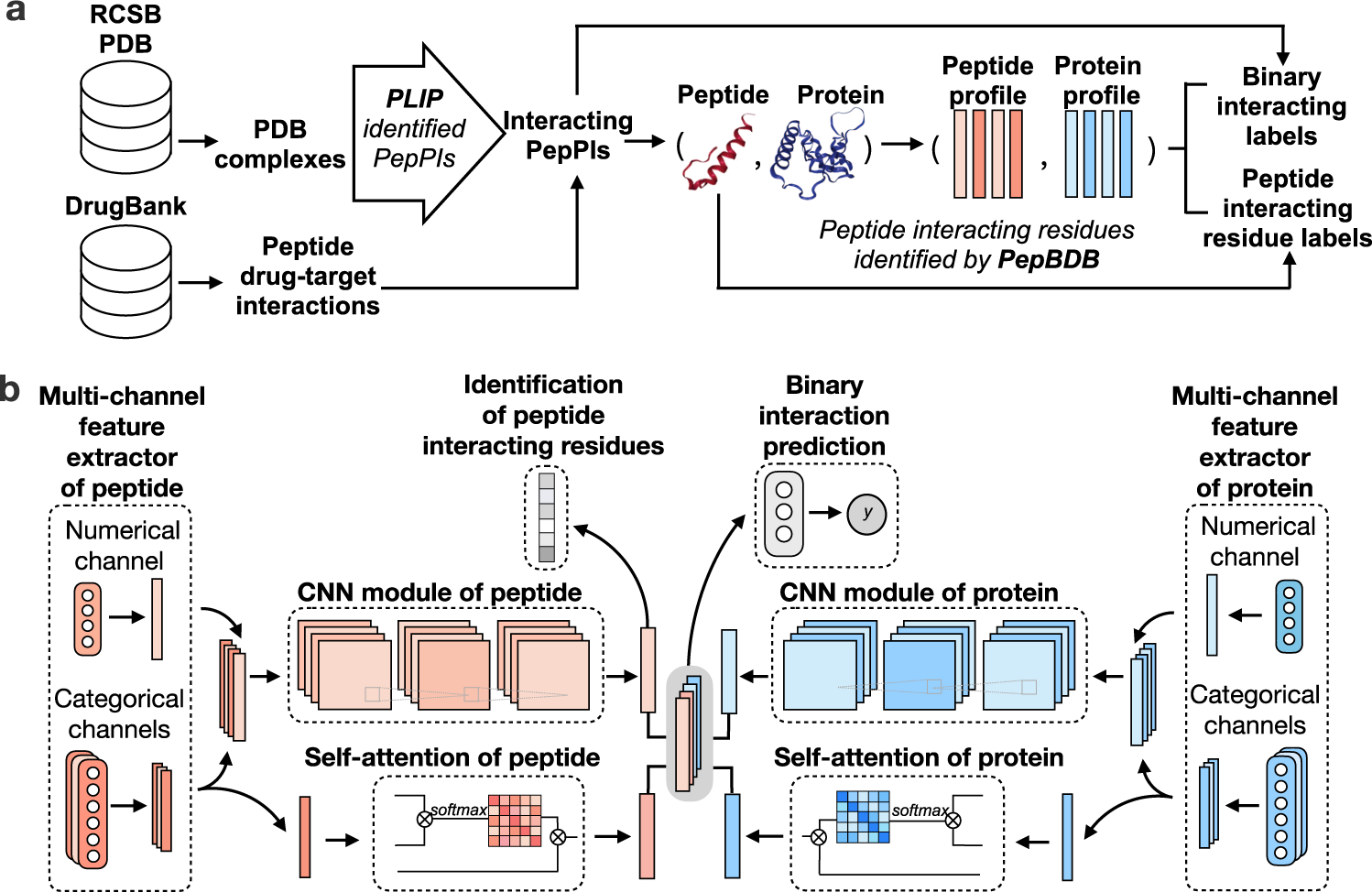
A deep-learning framework for multi-level peptide–protein interaction prediction | Nature Communications

In silico-prediction of protein–protein interactions network about MAPKs and PP2Cs reveals a novel docking site variants in Brachypodium distachyon | Scientific Reports

Antibody-protein interactions: benchmark datasets and prediction tools evaluation , Various Authors - Amazon.com

Characterizing Protein–Protein Interactions Using Mass Spectrometry: Challenges and Opportunities: Trends in Biotechnology